![PDF] Molecular Docking Using Chimera and Autodock Vina Software for Nonbioinformaticians | Semantic Scholar PDF] Molecular Docking Using Chimera and Autodock Vina Software for Nonbioinformaticians | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/52b25f8d3fb6c921297d2258ee9b36d05a2e33f6/20-Figure18-1.png)
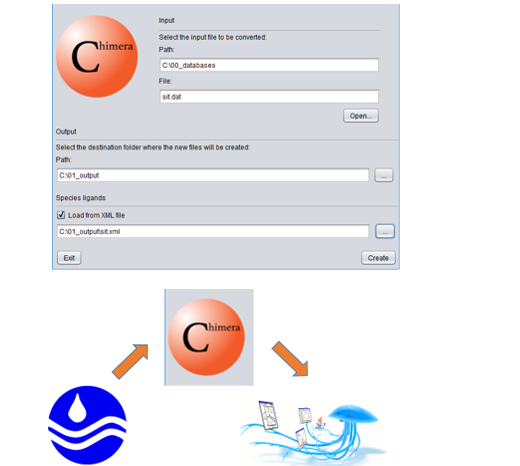
PDF] Molecular Docking Using Chimera and Autodock Vina Software for Nonbioinformaticians | Semantic Scholar
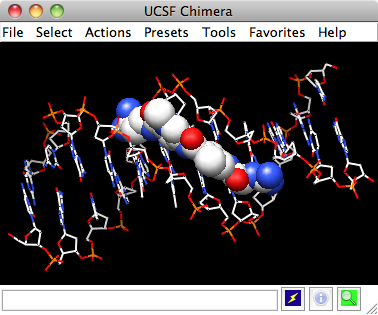
UCSF Chimera: Molecular Modeling Software for Visualization and Analysis of Molecular Structures - Available technology for licensing from the UCSF
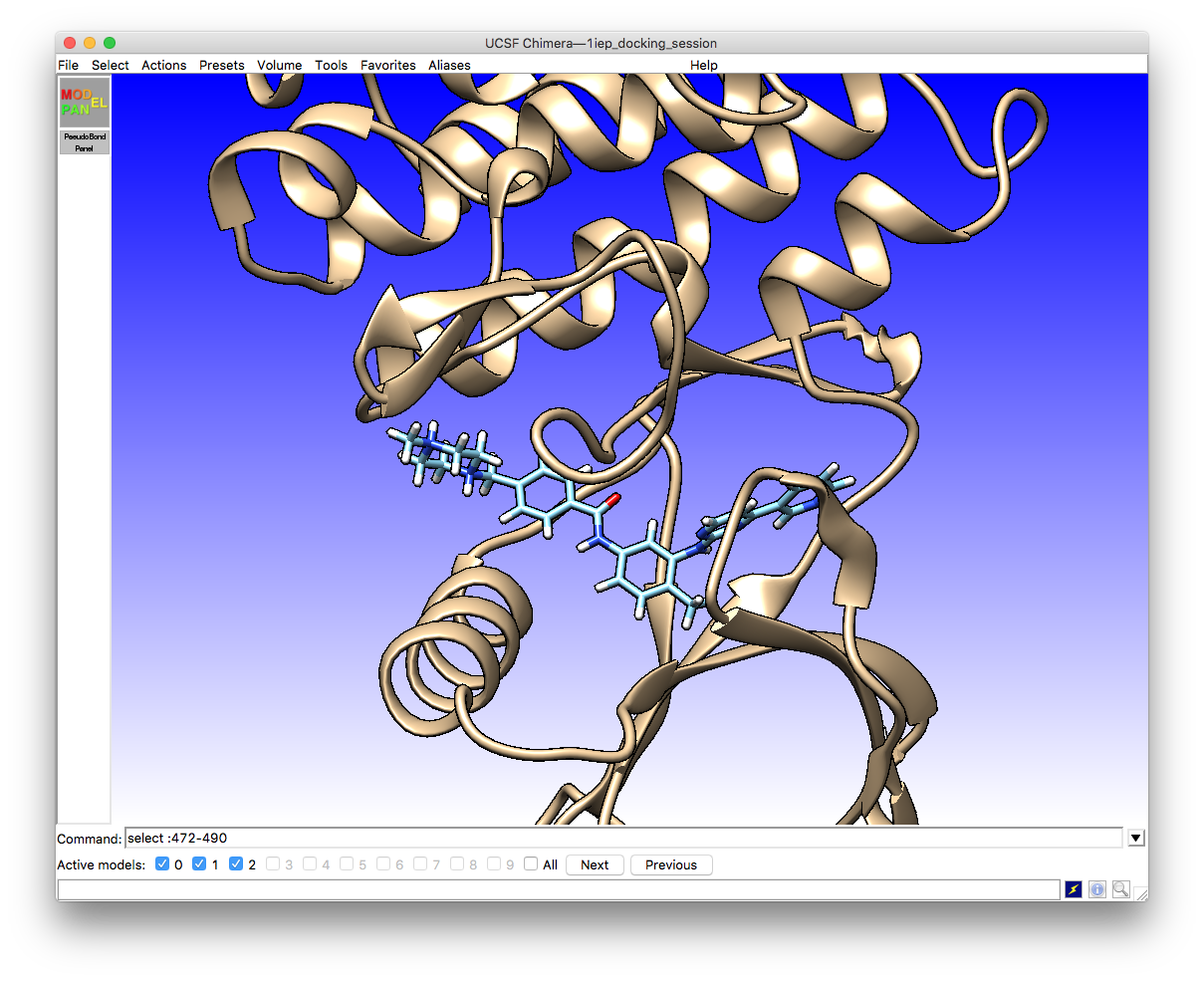
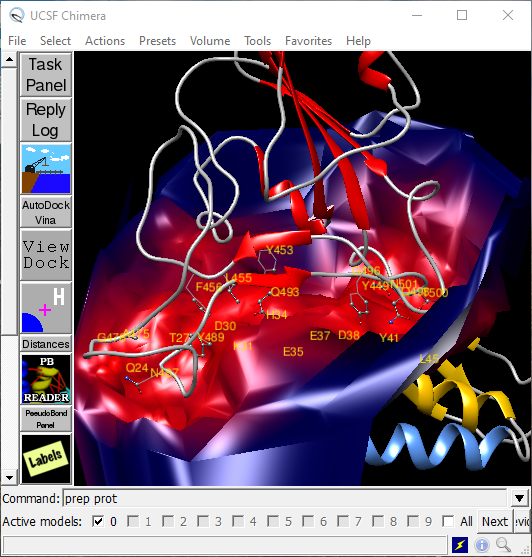
7. Protein-ligand docking with Chimera and Vina — ScotCHEM protein-ligand docking course documentation
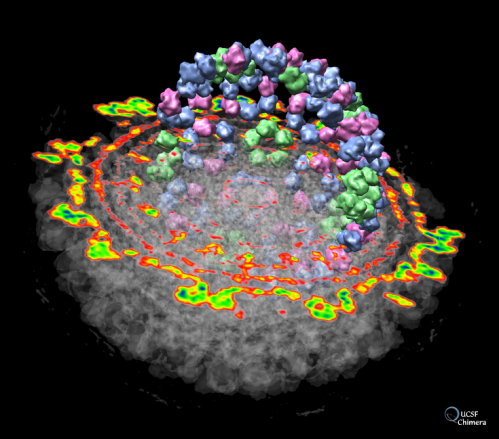
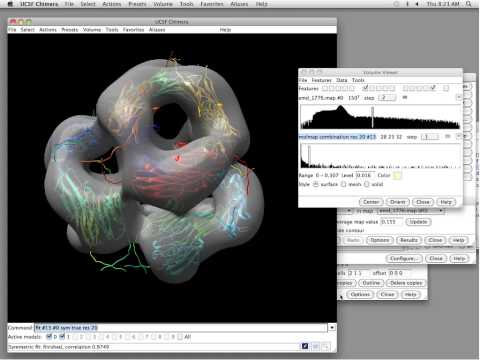
View coloured 3D map in Chimera using output from local resolution - Cryo-EM Data Processing - CryoSPARC Discuss





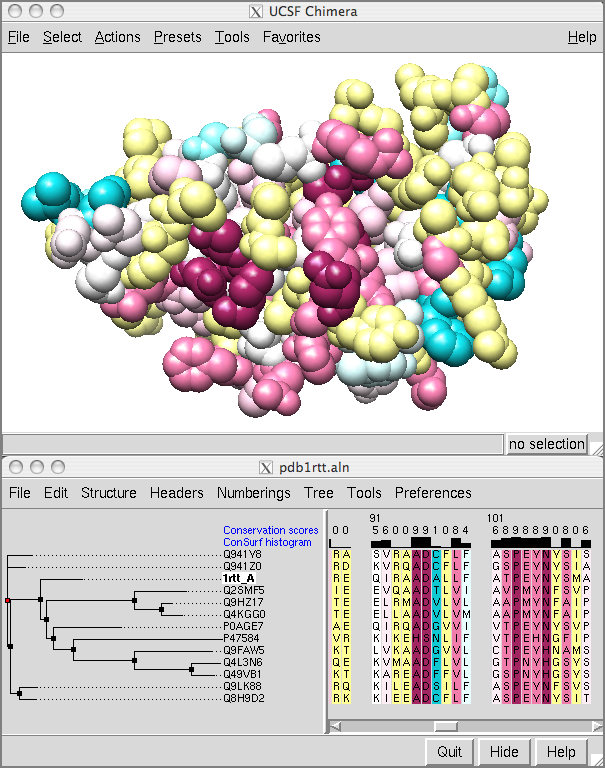

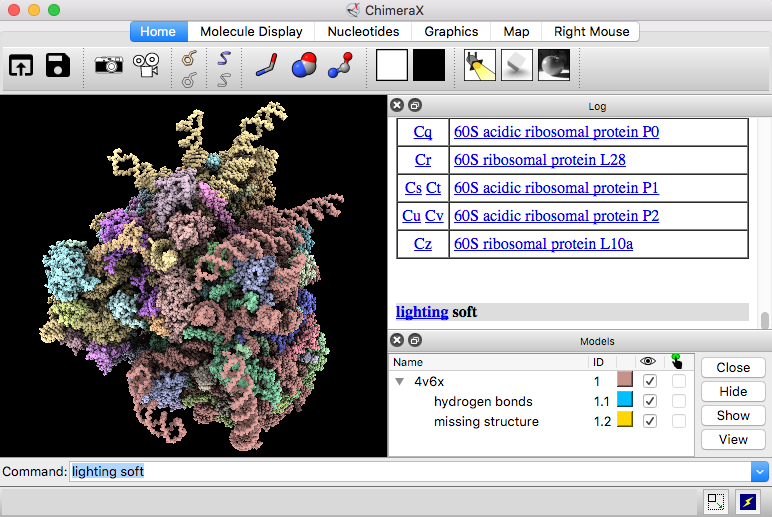
![ChimeraX] color cryo-EM local resolution – KPWu's group research site ChimeraX] color cryo-EM local resolution – KPWu's group research site](https://kpwulab.files.wordpress.com/2021/05/chimerax-surface_color.png)
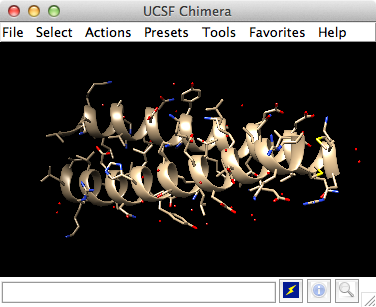
![Chimera] Coloring local resolution – KPWu's group research site Chimera] Coloring local resolution – KPWu's group research site](https://kpwulab.files.wordpress.com/2021/05/local_res.png?w=1024)